NEB Reaction Path Visualization¶
In this tutorial you will produce publication-quality figures from a Diels-Alder NEB calculation: a 1D energy profile, a 2D reaction valley landscape with GP surface [1] , structure galleries with four rendering backends, OCI-NEB peak overlays, and band evolution plots.
Reference
The 2D RMSD projection method used here is described in:
R. Goswami, “Two-dimensional RMSD projections for reaction path visualization and validation,” MethodsX, p. 103851, Mar. 2026, doi:10.1016/j.mex.2026.103851.
If you use these visualizations in your work, please cite the paper.
Prerequisites¶
# Core package with all rendering backends
pip install "rgpycrumbs[all]" xyzrender solvis-tools ovito
The tutorial data ships with the rgpycrumbs source tree at
docs/orgmode/tutorials/data/diels_alder/. It contains output from an
OCI-NEB calculation of the Diels-Alder [4+2] cycloaddition of
1,3-butadiene and ethylene (C6H10, 16 atoms, 22 images).
Step 1: Energy Profile¶
rgpycrumbs eon plt-neb \
--plot-type profile \
--input-dat-pattern "data/diels_alder/neb_*.dat" \
--input-path-pattern "data/diels_alder/neb_path_*.con" \
--con-file data/diels_alder/neb.con \
--sp-file data/diels_alder/sp.con \
-o profile.pdf
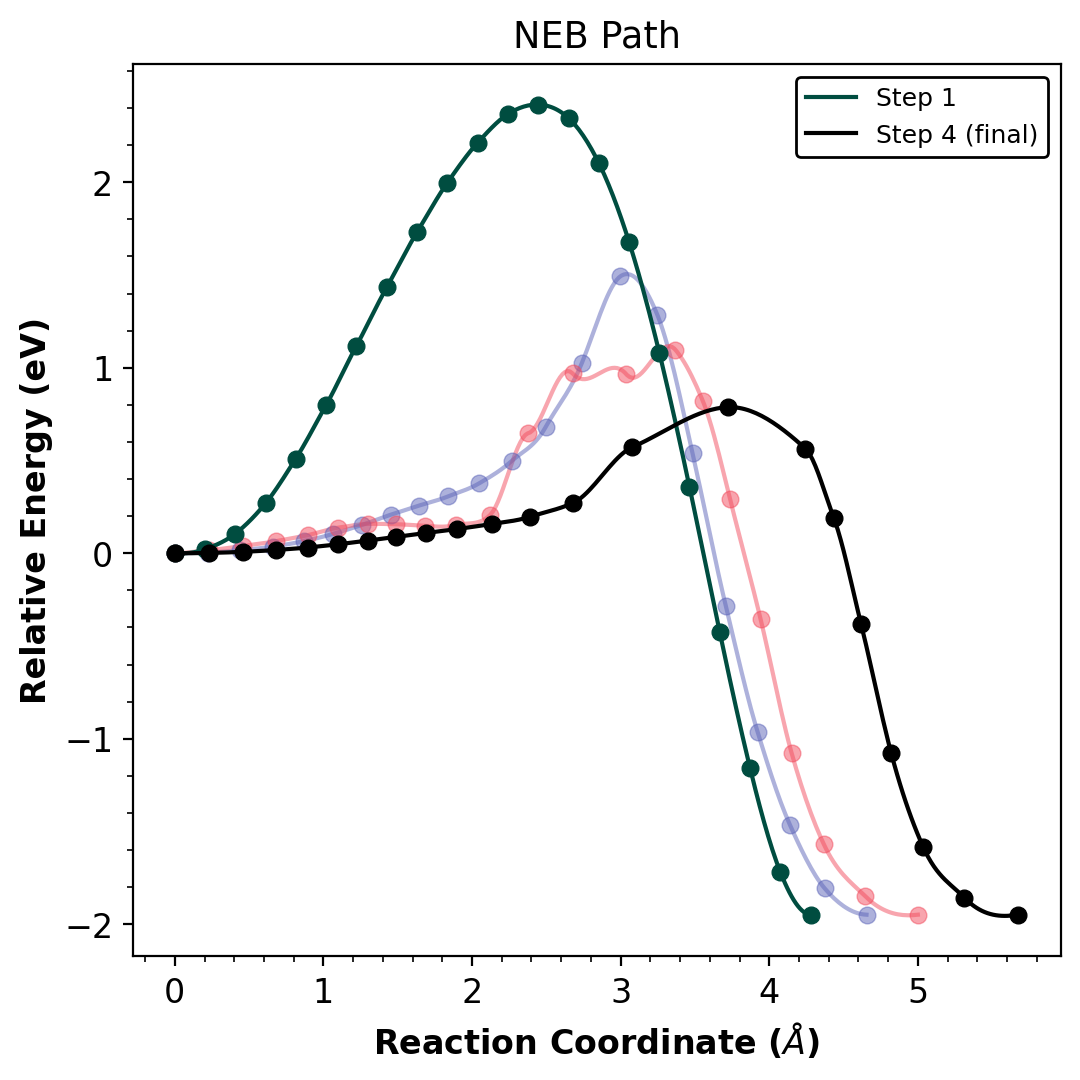
Multiple NEB optimization steps are overlaid with Hermite spline interpolation. The final converged barrier is ~0.8 eV.
Step 2: 2D Reaction Valley Landscape¶
Path overlay (no surface fit)¶
rgpycrumbs eon plt-neb \
--plot-type landscape \
--landscape-mode path \
--no-project-path \
--con-file data/diels_alder/neb.con \
--sp-file data/diels_alder/sp.con \
-o landscape_raw.pdf
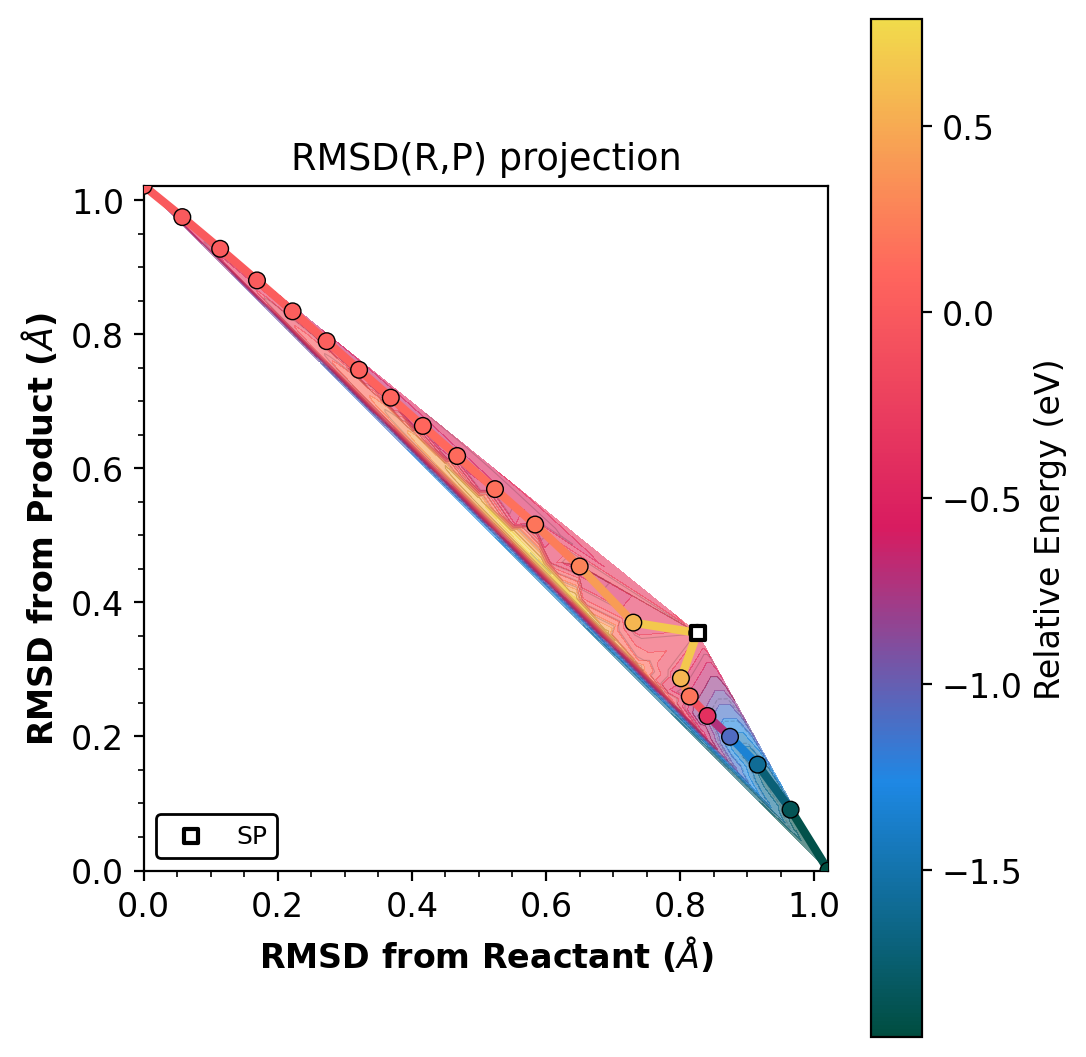
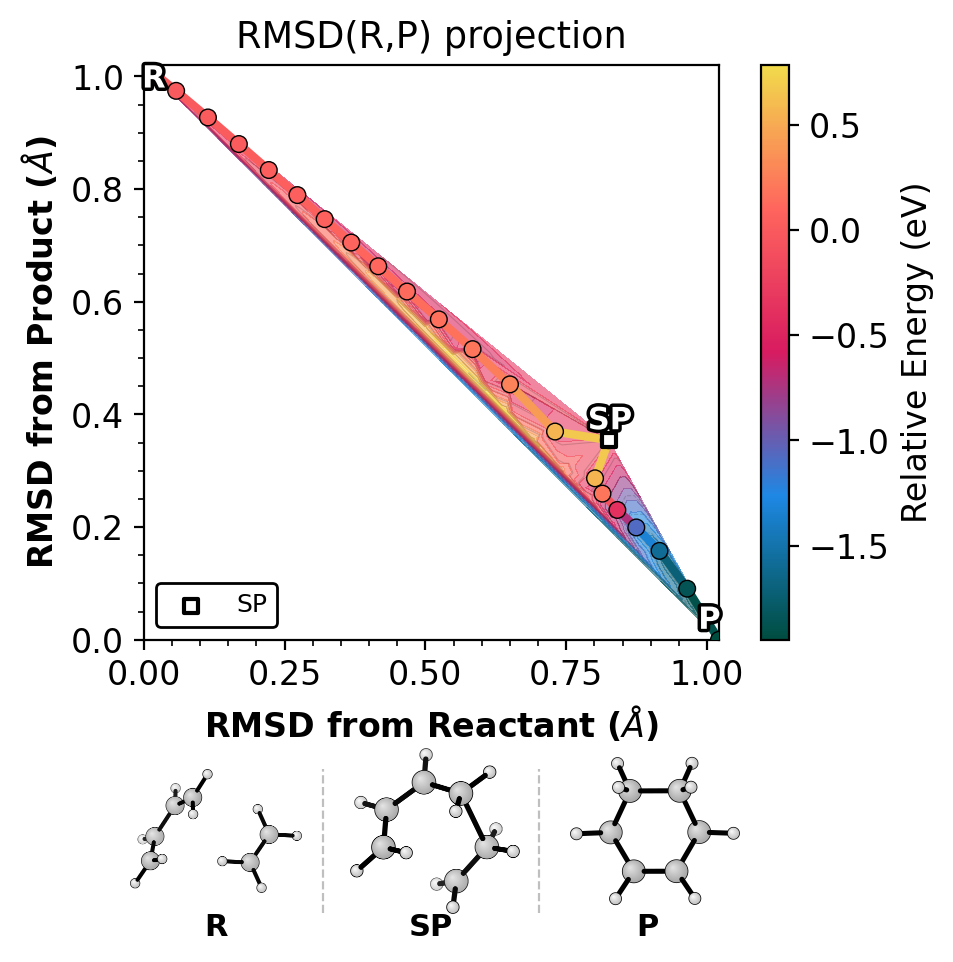
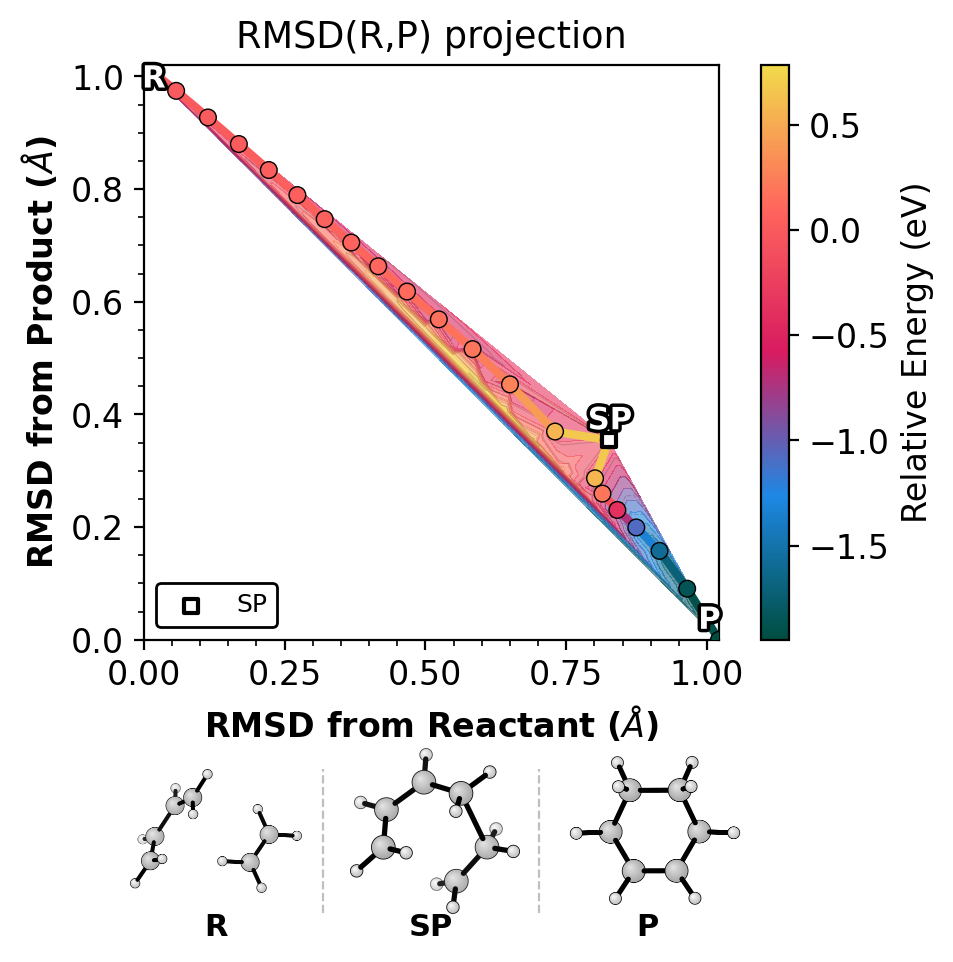
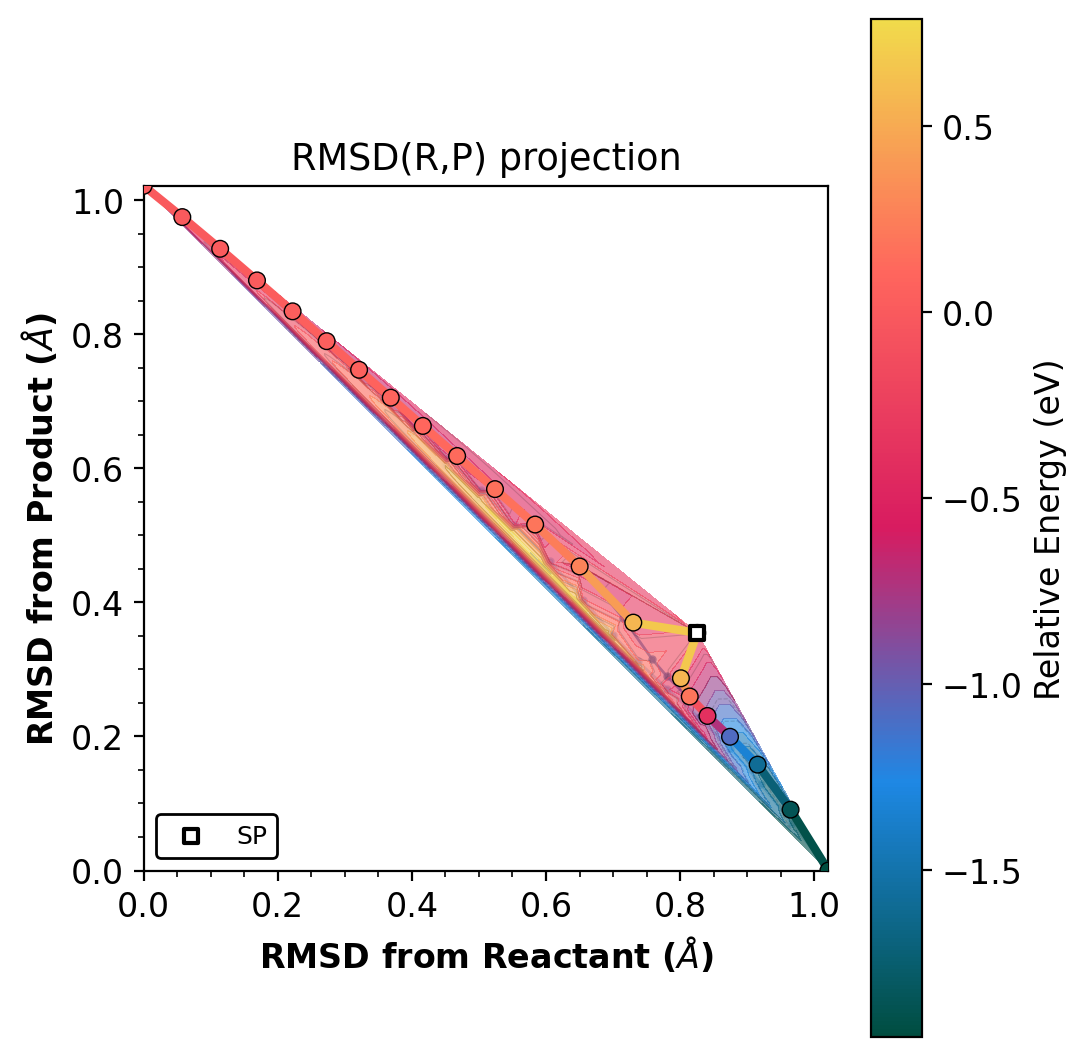
Shows the NEB path on raw RMSD(R,P) axes, colored by energy.
Projected (s, d) coordinates¶
rgpycrumbs eon plt-neb \
--plot-type landscape \
--landscape-mode path \
--project-path \
--con-file data/diels_alder/neb.con \
--sp-file data/diels_alder/sp.con \
-o landscape_projected.pdf
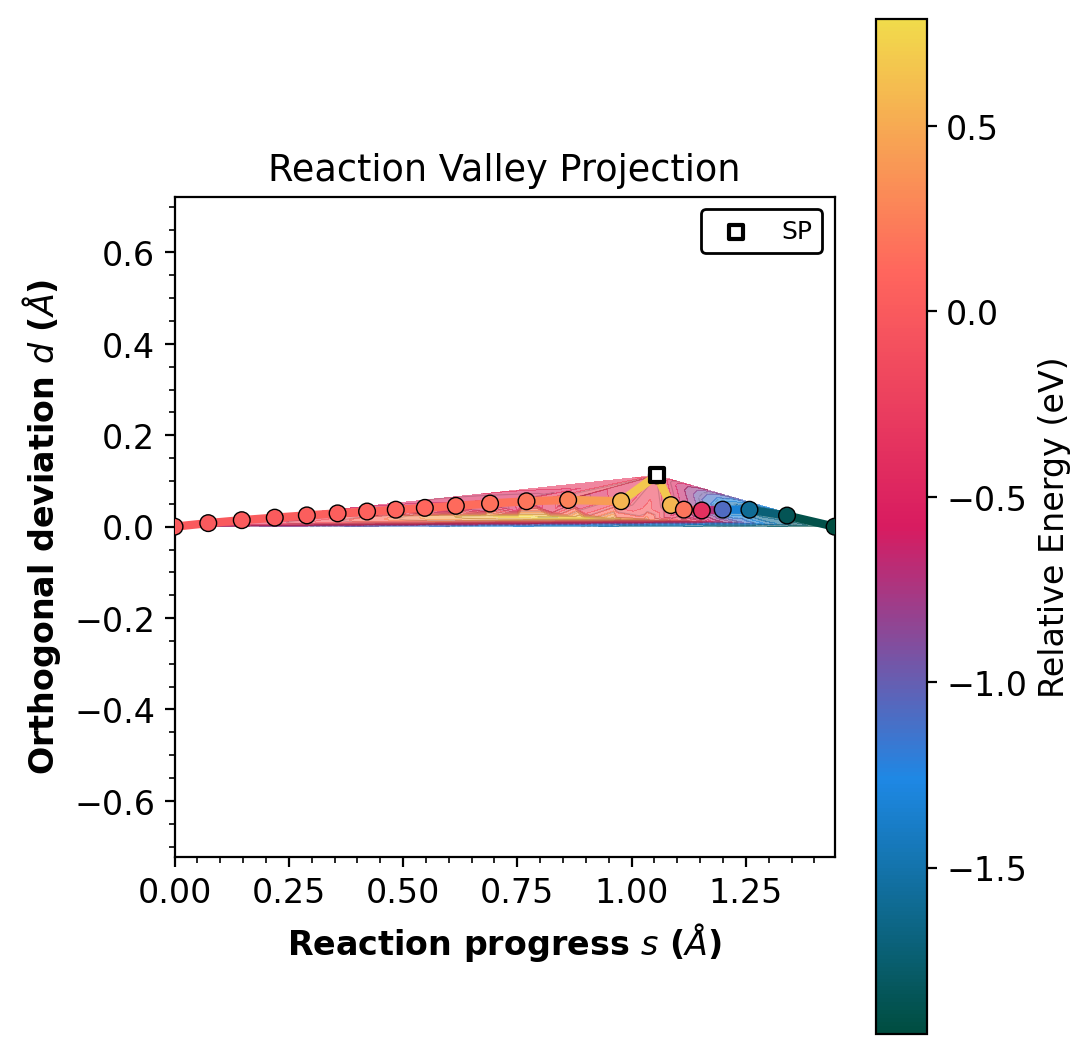
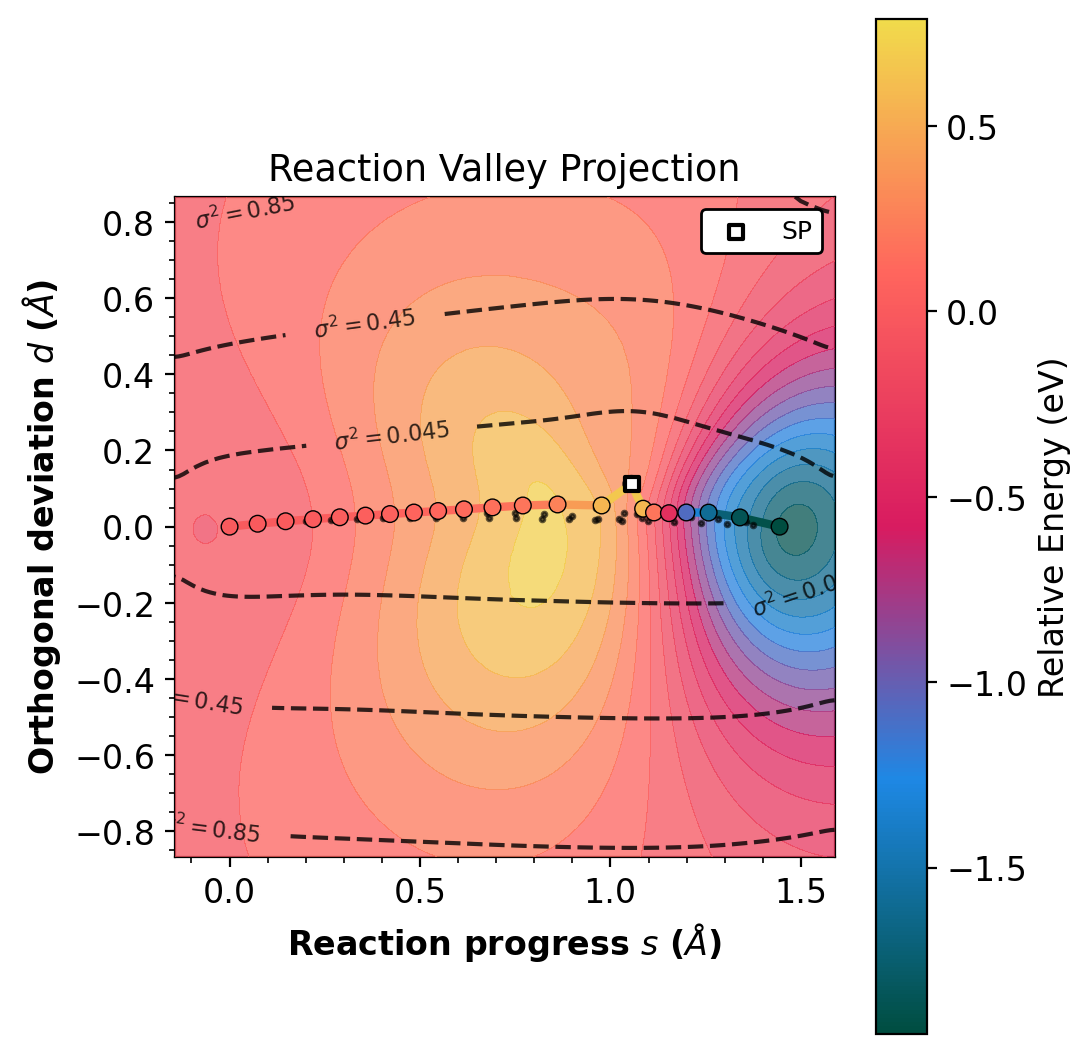
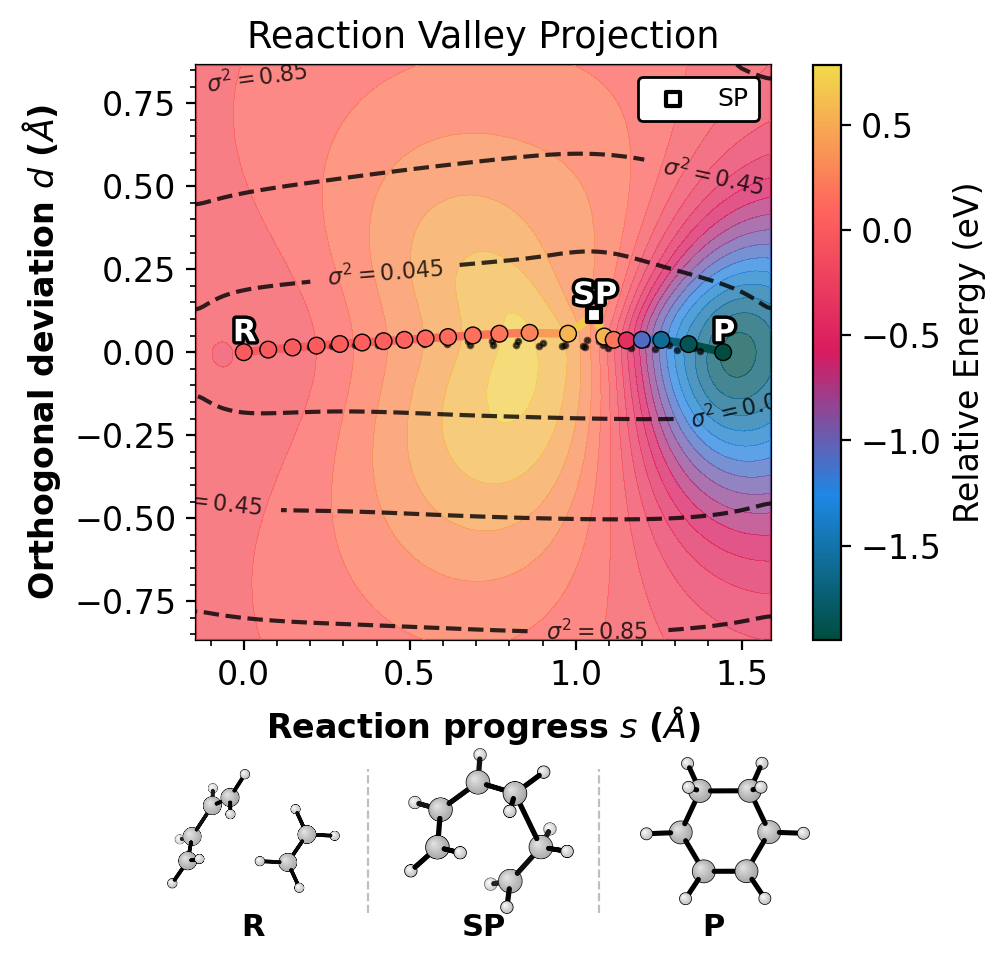
The --project-path flag rotates into reaction progress (s) and
orthogonal deviation (d). A path with d near 0 indicates a
concerted mechanism.
GP surface fit¶
rgpycrumbs eon plt-neb \
--plot-type landscape \
--landscape-mode surface \
--surface-type grad_matern \
--project-path \
--con-file data/diels_alder/neb.con \
--sp-file data/diels_alder/sp.con \
-o landscape_surface.pdf
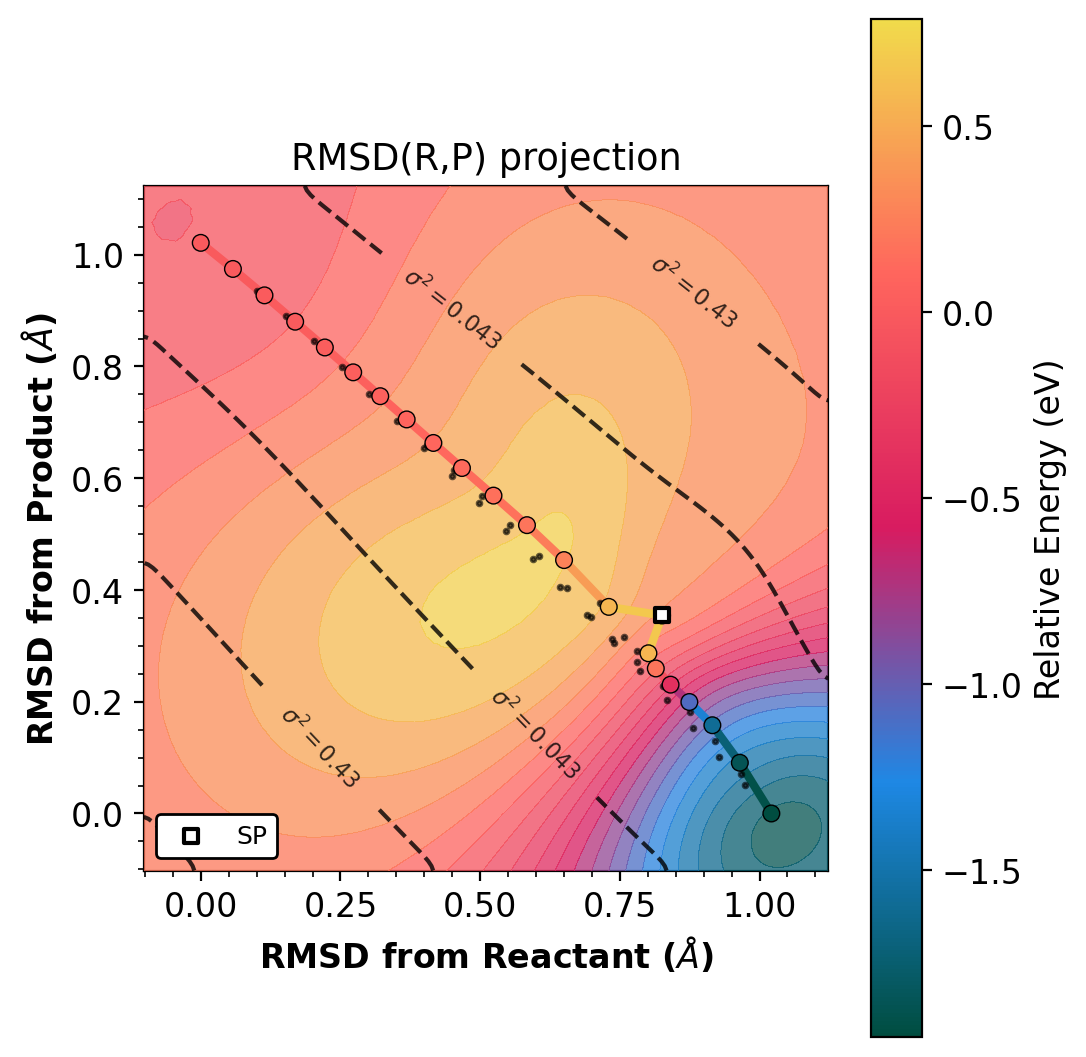
Left: projected (s,d). Right: raw RMSD(R,P). Gradient-enhanced Matern kernel GP surface with variance contours (dashed lines). Requires jax.
Step 3: Structure Galleries¶
Rendering backends¶
Four backends are available via --strip-renderer. Each wraps an
external visualization library:
xyzrender (default) – ball-and-stick with the
patonpreset, auto-orientation via PCAASE – space-filling spheres, built-in (no extra deps)
solvis – PyVista ball-and-stick with transparent RGBA backgrounds
OVITO Python – off-screen rendering with orthographic projection
See the Rendering Backends Guide for detailed feature comparison.
# xyzrender (default): ball-and-stick with paton preset
rgpycrumbs eon plt-neb \
--plot-type landscape --landscape-mode path --no-project-path \
--con-file data/diels_alder/neb.con --sp-file data/diels_alder/sp.con \
--plot-structures crit_points \
--strip-renderer xyzrender \
--strip-spacing 2.0 --strip-dividers \
-o gallery_xyzrender.pdf
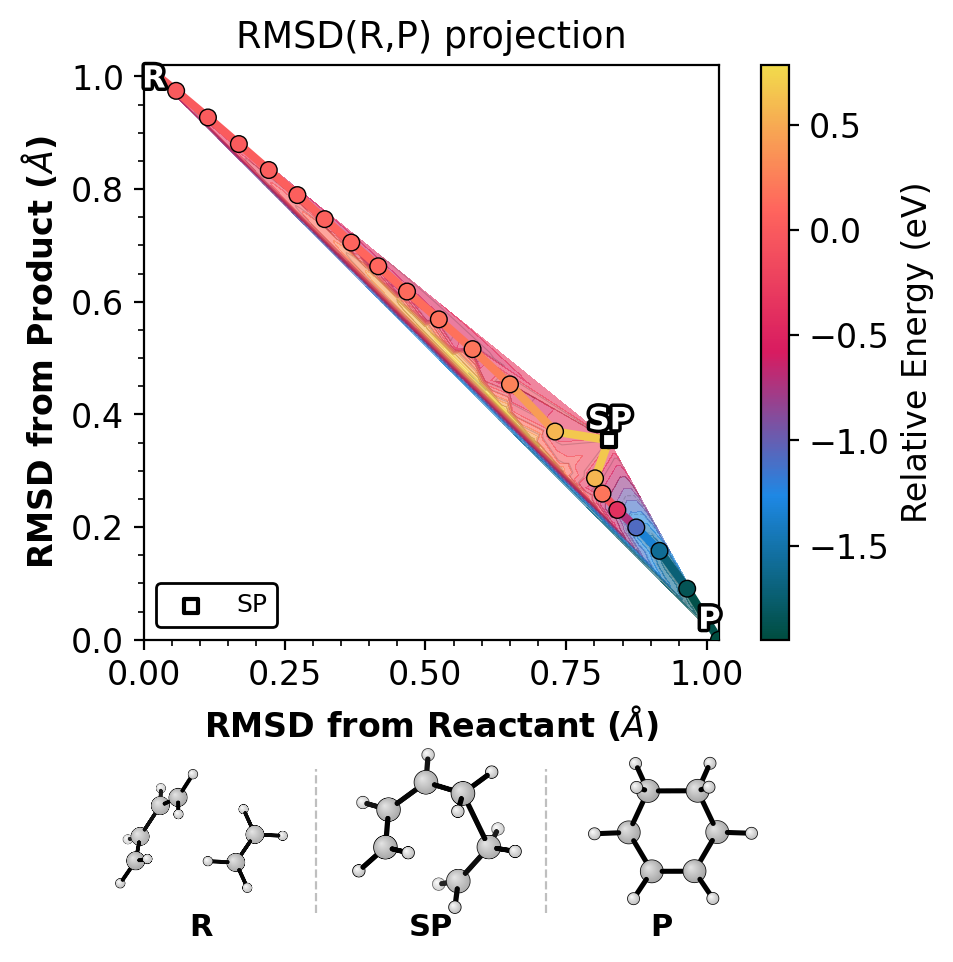
Left: xyzrender (ball-and-stick). Right: ASE (space-filling).
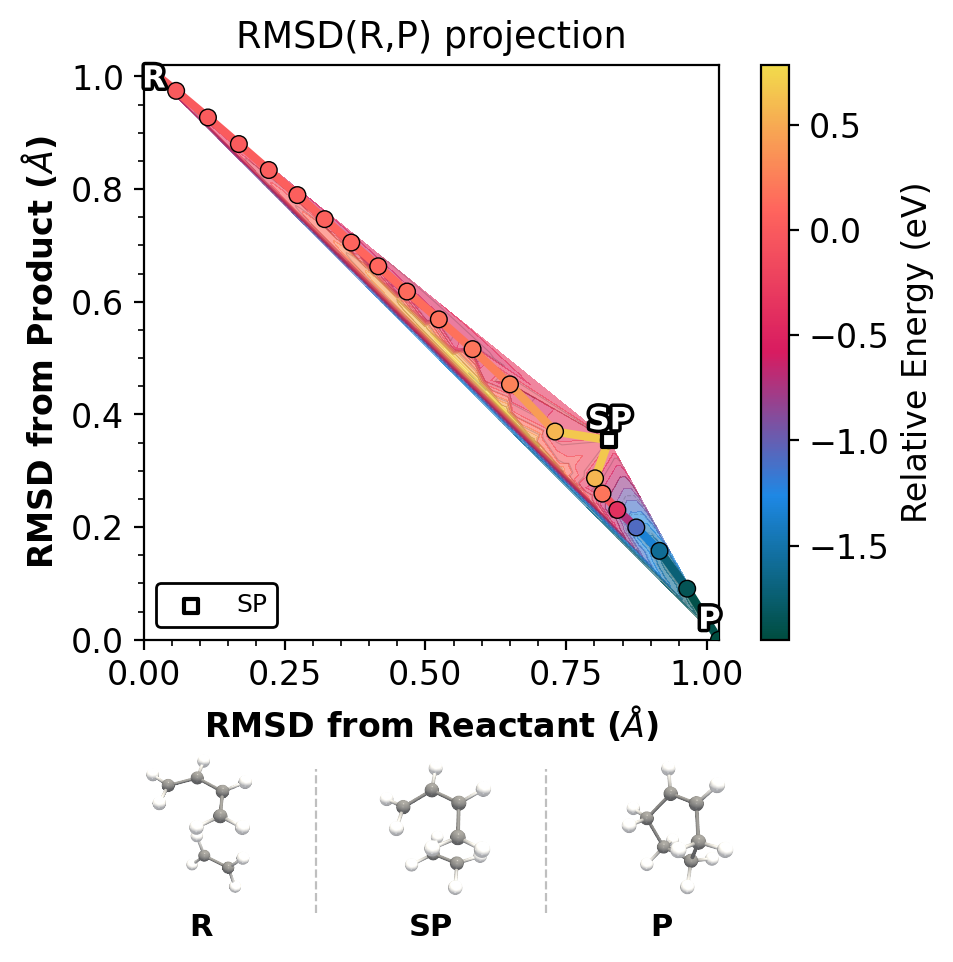
solvis: PyVista ball-and-stick with transparent background.
Viewing angle¶
The --rotation flag controls the viewing angle for all backends:
rgpycrumbs eon plt-neb ... --rotation "45x,30y,0z"
Perspective tilt¶
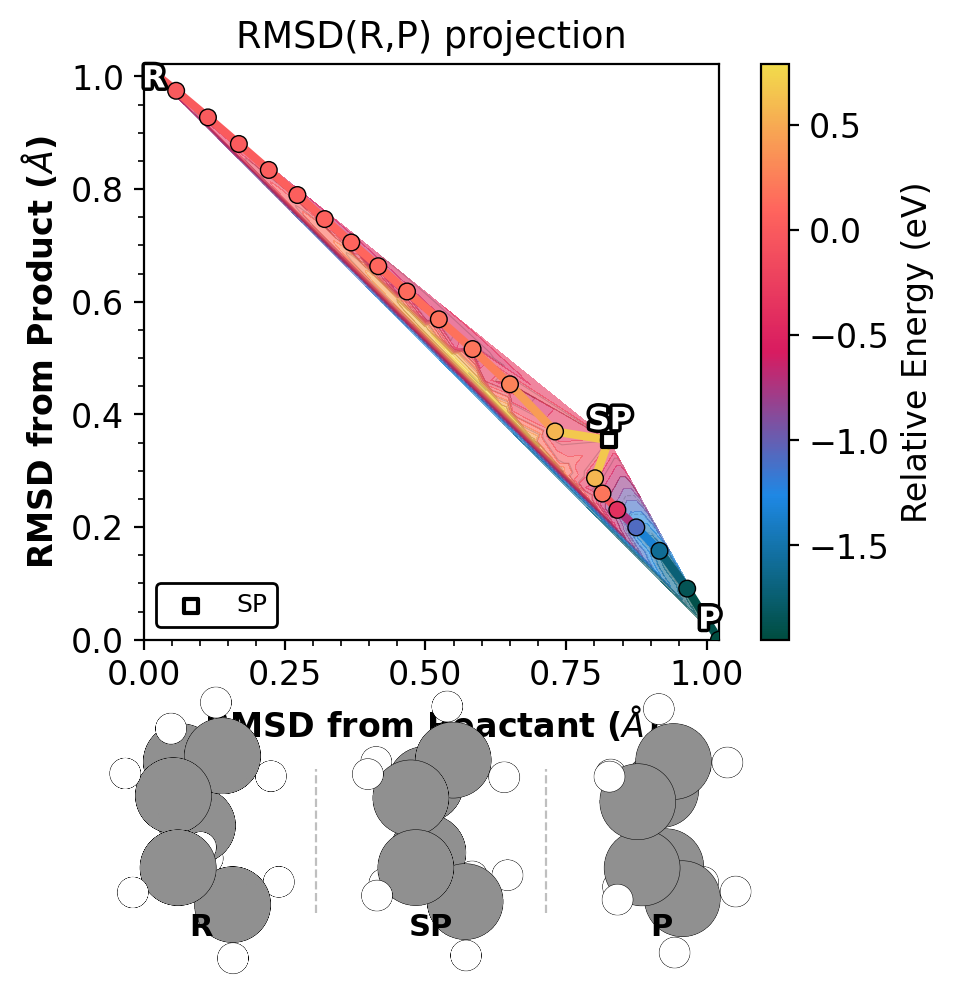
When atoms overlap in orthographic projection, use --perspective-tilt
to apply a small Rodrigues off-axis rotation that reveals hidden atoms:
rgpycrumbs eon plt-neb \
--plot-type landscape --landscape-mode path --no-project-path \
--con-file data/diels_alder/neb.con --sp-file data/diels_alder/sp.con \
--plot-structures crit_points \
--strip-dividers \
--perspective-tilt 8 \
-o gallery_tilted.pdf
Left: no tilt (H atoms hidden). Right: 8 degree tilt (H atoms revealed).
Step 4: OCI-NEB Peak Overlay¶
rgpycrumbs eon plt-neb \
--plot-type landscape --landscape-mode path --no-project-path \
--con-file data/diels_alder/neb.con --sp-file data/diels_alder/sp.con \
--mmf-peaks --peak-dir data/diels_alder/ \
-o mmf_peaks.pdf
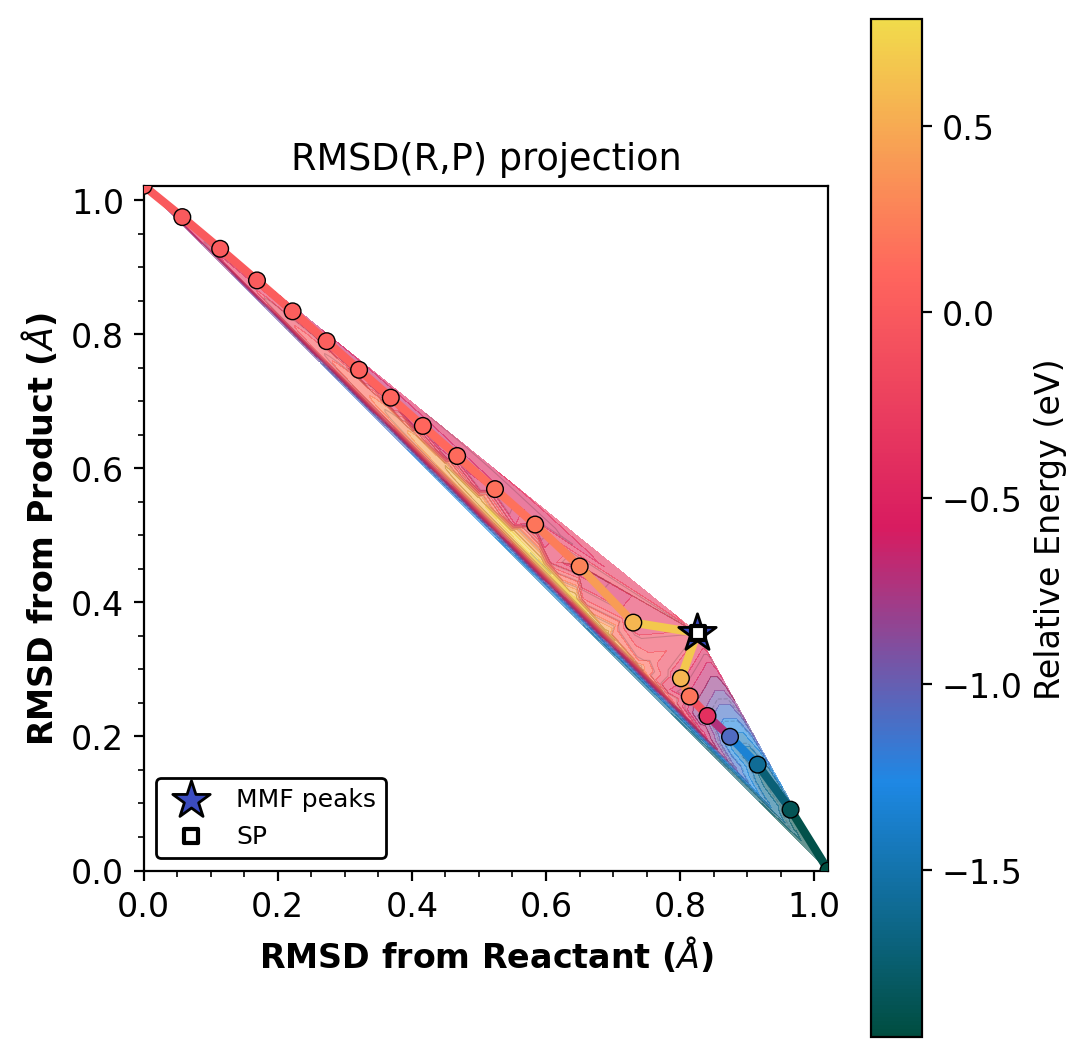
Star markers show where OCI-NEB dimer refinement placed the saddle.
Step 5: Band Evolution¶
rgpycrumbs eon plt-neb \
--plot-type landscape --landscape-mode path --no-project-path \
--con-file data/diels_alder/neb.con \
--show-evolution \
-o evolution.pdf
Older bands are drawn with lower opacity.
Step 6: Combined Figure¶
The full figure with 2D GP surface, xyzrender structure strip, and MMF peak overlay:
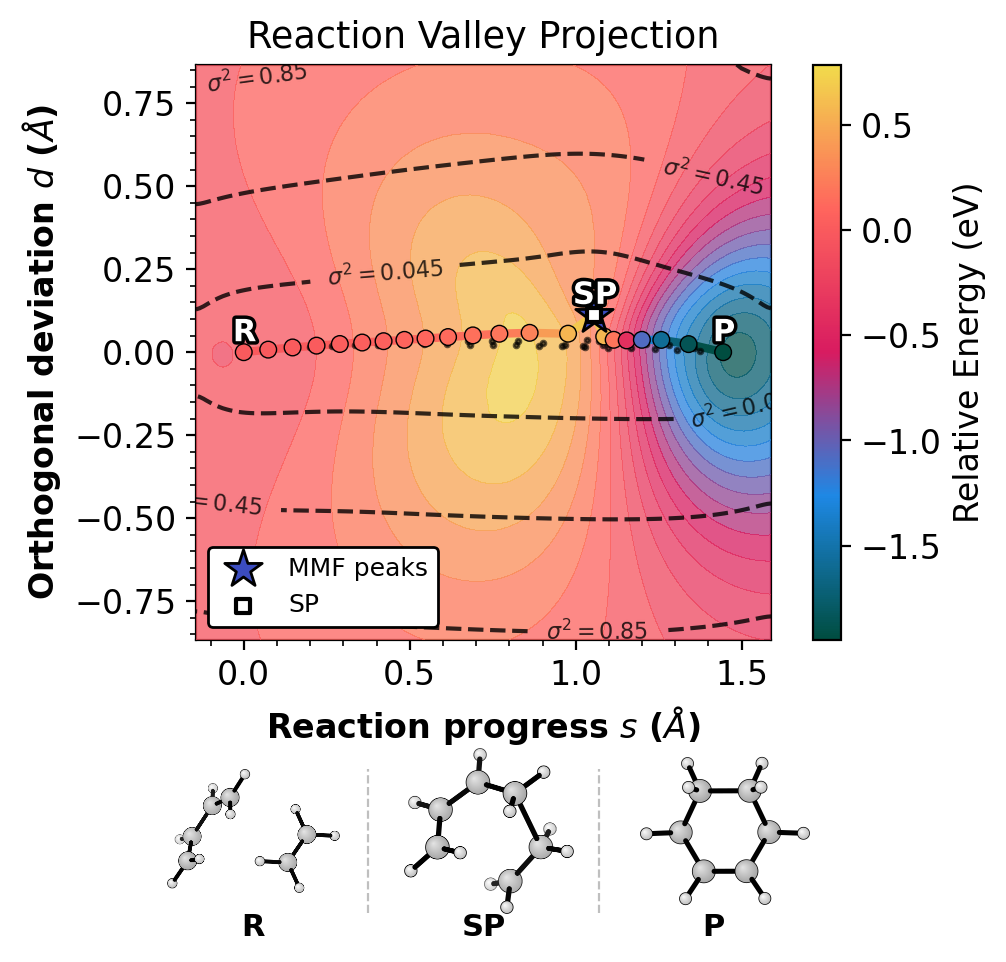
rgpycrumbs eon plt-neb \
--plot-type landscape --landscape-mode surface \
--surface-type grad_matern --project-path \
--con-file data/diels_alder/neb.con --sp-file data/diels_alder/sp.con \
--mmf-peaks --peak-dir data/diels_alder/ \
--plot-structures crit_points \
--strip-renderer xyzrender \
--strip-spacing 2.0 --strip-dividers \
-o combined.pdf
Summary¶
Feature |
Flag |
Notes |
|---|---|---|
Energy profile |
|
Hermite spline |
Raw RMSD landscape |
|
RMSD(R) vs RMSD(P) |
Reaction valley |
|
(s, d) coordinates |
GP surface |
|
Needs jax |
Structure strip |
|
R, SP, P |
xyzrender |
|
Default, ball-and-stick |
ASE |
|
Space-filling |
solvis |
|
PyVista, transparent bg |
ovito |
|
OVITO off-screen |
Custom rotation |
|
All backends |
Perspective tilt |
|
Reveals hidden atoms |
Spacing/dividers |
|
|
MMF peaks |
|
OCI-NEB |
Band evolution |
|
Fading iterations |
See Also¶
Goswami, “Two-dimensional RMSD projections for reaction path visualization and validation,” MethodsX, p. 103851, Mar. 2026, doi:10.1016/j.mex.2026.103851